Zatsepin Laboratory
We run several projects devoted to RNA Biology and RNA Therapeutics in vitro and in vivo
1) RNA therapeutics based on modified antisense/siRNA/sgRNA with improved efficacy, biodistribution and stability in vivo.
We are working on the development of RNA-based therapeutics:
– novel antisense oligonucleotides that target RNA splicing to treat hereditary diseases in vitro and in vivo;
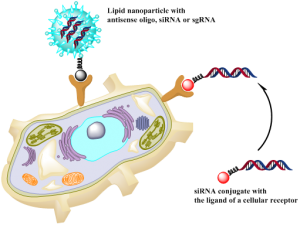
– caged siRNA and hybrid nanoparticles to target liver diseases accompanied by inflammation;
– siRNA conjugates to validate targets in the liver;
– chemically modified guide RNA for improved selectivity of genome editing
Joint projects with Prof. Gorin (CPQM, Skoltech) on novel functional nanomaterials for RNA delivery in vitro and in vivo.
2) Validation of RNA helicases and lncRNA as therapeutic targets in vitro and in vivo
– Role of DDX3 helicase in the translation process and as a therapeutic target. Varied level of DDX3 allows us to reveal target mRNAs for DDX3 RNA helicase, hence, define and describe the exact role of DDX3 in translation in the mouse liver.
– Role of LL35 lncRNA in the liver. We found that functional analog of the ncRNA DEANR1 in mouse – LL35 ncRNA is involved in the regulation of transcription. Transcriptome and proteome analysis revealed that LL35 can affect metabolic pathways in liver cells.
| Principal Investigator – Timofei Zatsepin |  |
Team: |
|
||
| Olga Sergeeva | Tatiana Abakumova | Nataliya Logvina | |
| Svetlana Dukova | |||
PhD Students: |
|||
| Denis Melnik | Evgeniya Shcherbinina | ||
Alumni: |
|||
| Renata Ialchina, MS student in 2017-18; current: PhD student University of Copenhagen, Denmark | Philipp Sergeev, MS student in 2018-19; current: PhD student in Institute for Molecular Medicine, Finland | Mariia Nazarova, MS student in 2018-19; current: PhD student in The Research Institute of Molecular Pathology, Austria | Svetlana Korinfskaya, MS student in 2017-18; current: PhD student in Cincinnati Children’s Hospital Medical Center, USA |
| Dr. Ilya Aparin, junior researcher in 2018-19 – current: postdoc in University of Strasbourg, France | Dominique Leboeuf, PhD student 2017-20; current: scientist in Beam Therapeutics, USA | Ilia Kopnin, MS student in 2019-20; current: PhD student at KU Leuven, Belgium | Aleksandr Shikalov, MS student in 2019-20; |
- Silver(I)-mediated base pairing in DNA involving the artificial nucleobase 7,8-dihydro-8-oxo-1,N6-ethenoadenine. Schönratha V, Tsvetkov V, Barceló-Oliverd M, Hebenbrocka M, Zatsepin T, Aralov F, Müllera J. Journal of Inorganic Biochemistry. https://doi.org/10.1016/j.jinorgbio.2021.111369
- Novel Lipid-Oligonucleotide Conjugates Containing Long-Chain Sulfonyl Phosphoramidate Groups: Synthesis and Biological Properties Derzhalova A, Markov O, FokinaO, Shiohama Y, Zatsepin T, Fujii M, Zenkova M, Stetsenko D Appl. Sci. 2021, 11(3), 1174; https://doi.org/10.3390/app11031174
- Multifunctional nanostructured drug delivery carriers for cancer therapy: Multimodal imaging and ultrasound-induced drug release Novoselova M, German S, Abakumova T, Perevoschikov S, Sergeeva O, Nesterchuk M, Efimova O, Petrov K, Chernyshev V, Zatsepin T, Gorin D. Colloids Surf B Biointerfaces. 2021 Jan 20;200:111576. doi: 10.1016/j.colsurfb.2021.111576
- Genomic DNA i-motifs as fast sensors responsive to near-physiological pH microchanges. Turaev A, Isaakova E, Severov V, Bogomazova A, Zatsepin T, Sardushkin M, Aralov A, Lagarkova M, Pozmogova G, Varizhuk A. Biosensors and Bioelectronics
Volume 175, 1 March 2021, 112864
- Toehold-Mediated Selective Assembly of Compact Discrete DNA Nanostructures. V.A.Brylev, A.V.Ustinov, V.B.Tsvetkov, N.A.Barinov, I.O.Aparin, K.A.Sapozhnikova, Y.Y.Berlina, E.A.Kokin, D.V.Klinov, T.S.Zatsepin, V.A.Korshun. Langmuir. 2020 Dec 15;36(49):15119-15127. doi: 10.1021/acs.langmuir.0c02696.
- Multispectral sensing of biological liquids with hollow-core microstructured optical fibres. Ermatov T, Noskov R, Machnev A, Gnusov I, Аtkin V, Lazareva E, German S, Kosolobov S, Zatsepin T, Sergeeva O, Skibina J, Ginzburg P, Tuchin V, Lagoudakis P, Gorin D. Light: Science & Applications, 9, 173 (2020)
- EEF2 depletion by RNAi in mouse liver leads to mTOR-independent translational upregulation of ribosomal genes. Gerashchenko M, Nesterchuk M, Smekalova E, Paulo J, Kowalski P, Gygi S, Rogorad B, Zatsepin T, Anderson D, Gladyshev V, Kotelianski V. Sci. Rep., 10 (1), 15473 (2020).
- Comparative Analysis of Long Noncoding RNA Expression in Human Hepatocyte Cell Lines and Liver. Burenina O, Zatsepin T, Kim E, Metelin A, Skvortsov D, Rubtsova M, Dontsova O. Biochem. Biophys. Volume 493, pages181–184(2020)
- Panel of potential lncRNA biomarkers can distinguish various types of liver malignant and benign tumors Burenina O, Lazarevich N, Kustova I, Shavochkina D, Moroz E, Kudashkin N, Patyutko Y, Metelin A, Kim E, Skvortsov D, Zatsepin T, Rubtsova M, Dontsova O. J Cancer Res Clin Oncol. 2020 Sep 12. doi: 10.1007/s00432-020-03378-5. PMID: 32918630 DOI: 10.1007/s00432-020-03378-5
- Upregulation of Mcl-1S Causes Cell-Cycle Perturbations and DNA Damage Accumulation. Streletskaia A, Senichkin A, Prikazchikova T, Zatsepin T Zhivotovsky B, Kopeina G Front. Cell Dev. Biol. 25.09.2020 https://doi.org/10.3389/fcell.2020.543066
- Red light-triggered photoreduction on a nucleic acid template Dutta S, Rühle J, Schikora M, Deussner-Helfmann N, Heilemann M, Zatsepin T, Duchstein P, Zahn D, Knör G, Mokhir A. Chem Commun. 30.07.2020. doi: 10.1039/d0cc03086d.
- Murine Long Noncoding RNA Morrbid Contributes in the Regulation of NRAS Splicing in Hepatocytes In Vitro. Fefilova A, Melnikov P, Prikazchikova P, Abakumova T, Kurochkin I, Mazin P, Ziganshin R, Sergeeva O, Zatsepin T. Int J Mol Sci. 05.08.2020;21(16):E5605. doi: 10.3390/ijms21165605.PMID: 32764370 DOI: 10.3390/ijms21165605
- Translation at first sight: the influence of leading codons. Osterman I, Chervontseva Z , Evfratov S, Sorokina A, Rodin V, Rubtsova R, Komarova E, ZatsepinT, Kabilov M, Bogdanov A, Gelfand M, Dontsova O, Sergiev P. 09.07.2020;48(12):6931-6942. doi: 10.1093/nar/gkaa430.
- Optical clearing for photoacoustic lympho- and angiography beyond conventional depth limit in vivo. Novoselova M, Abakumova T, Khlebtsov B, Zatsepin T, Lazareva E, Tuchin V, Zharov V, Gorin D, Galanzha E Photoacoustics. 17.06.2020;20:100186. doi: 10.1016/j.pacs.2020.100186. eCollection 2020 Dec.
- The Arg/N-Degron Pathway-A Potential Running Back in Fine-Tuning the Inflammatory Response? Leboeuf D, Pyatkov M, Zatsepin T, Piatkov K. Biomolecules. 14.06.2020;10(6):903. doi: 10.3390/biom10060903. PMID: 32545869 PMCID: PMC7356051 DOI: 10.3390/biom10060903
- Excimer-FRET Cascade in Dual DNA Probes: Open Access to Large Stokes Shift, Enhanced Acceptor Light up, and Robust RNA Sensing. Aparin I, Sergeeva O, Mishin A, Khaydukov E, Korshun V, Zatsepin T. Anal Chem. 05.05.2020. doi: 10.1021/acs.analchem.0c00270.
- Modification of Adenosine196 by Mettl3 Methyltransferase in the 5′-External Transcribed Spacer of 47S Pre-rRNA Affects rRNA Maturation. Sergeeva O, Sergeev P, Melnikov P, Prikazchikova T, Dontsova O, Zatsepin T. Cells. 24.04.2020;9(4). pii: E1061. doi: 10.3390/cells9041061.
- Influence of the spacer region between the Shine-Dalgarno box and the start codon for fine-tuning of the translation efficiency in Escherichia coli. Komarova E, Chervontseva Z, Osterman I, Evfratov S, Rubtsova M, Zatsepin T, Semashko T, Kostryukova E, Bogdanov A, Gelfand M, Dontsova O, Sergiev P. Microb Biotechnol. 23.03.2020. doi: 10.1111/1751-7915.13561.
- Short Duplex Module Coupled to G-Quadruplexes Increases Fluorescence of Synthetic GFP Chromophore Analogues Zaitseva S, Baleeva N, Zatsepin T, Myasnyanko I, Turaev M, Pozmogova G, Khrulev A, Varizhuk A, Baranov A, Aralov A. Sensors. 09.02.2020 , DOI: 10.3390/s20030915
-
Downregulation of the Arg/N-degron Pathway Sensitizes Cancer Cells to Chemotherapy in vivo Leboeuf D, Abakumova T, Prikazchikova T, Rhym L, Anderson D, Zatsepin T, Piatkov K. Mol. Ther. https://doi.org/10.1016/j.ymthe.2020.01.021
-
Biodegradable Polymeric Multilayer Capsules for Therapy of Lung Cancer. Novoselova M, Loh H, Trushina D, Ketkar A, Abakumova T, Zatsepin T, Kakran M, Brzozowska A, Lau H, Gorin D, Antipina M, Brichkina A. ACS Appl Mater Interfaces. 05.02.2020; 12(5):5610-5623. doi: 10.1021/acsami.9b21381.
- RT-qPCR Detection of Low-Copy HIV RNA with Yin-Yang Probes. Kireev D, Farzan V, Shipulin G, Korshun V, Zatsepin T. Methods Mol Biol. 2020; 2063:27-35. doi: 10.1007/978-1-0716-0138-9_3.
- DNA detection with dye-labeled oligonucleotides by surface enhanced Raman spectroscopy. Mendeleev Comm. Eremina O, Zatsepin T, Farzan V, Veselova I, Zvereva M, 30 (1), 18-21 (2020).
- In Vivo RNAi-Mediated eIF3m Knockdown Affects Ribosome Biogenesis and Transcription but Has Limited Impact on mRNA-Specific Translation. Smekalova E, Gerashchenko M, O’Connor P, Whittaker C, Kauffman K, Fefilova A, Zatsepin T, Bogorad R, Baranov P, Langer R, Gladyshev V, Anderson D, Koteliansky V. Mol Ther Nucleic Acids. 18.11.2019; 19:252-266. doi: 10.1016/j.omtn
- NHEJ pathway is involved in post-integrational DNA repair due to Ku70 binding to HIV-1 integrase. Knyazhanskaya E, Anisenko A, Shadrina O, Kalinina A, Zatsepin T, Zalevsky A, Mazurov D, Gottikh M; Retrovirology. 06.11.2019;16(1):30. doi: 10.1186/s12977-019-0492-z.
- Focused ultrasound-mediated fluorescence of composite microcapsules loaded with magnetite nanoparticles: In vitro and in vivo study. Novoselova M, Voronin D, Abakumova T, Demina P, Petrov A, Petrov V, Zatsepin T, Sukhorukov G, Gorin D. Colloids Surf B Biointerfaces. 01.09.2019;181:680-687. doi: 10.1016/j.colsurfb.2019.06.025.
- Silver(I)-mediated base pairing in parallel-stranded DNA involving the luminescent cytosine analog 1,3-diaza-2-oxophenoxazine. Schönrath I, Tsvetkov V, Zatsepin T, Aralov A, Müller J. J. Biol. Inorg. Chem., 24.08.2019;24(5):693-702. doi: 10.1007/s00775-019-01682-1
- Direct injection of SWCNTs into liquid after supercritical nitrogen treatment. Kalachikova P, Goldt A, Khabushev E, Eremin T, Ustinovich K, Grebenko A, Parenago O, Zatsepin T, Pokrovskiy O, Obraztsova E, Nasibulin A. Carbon, 2019. https://doi.org/10.1016/j.carbon.2019.06.003
- Robust technique for dispersion of single-walled carbon nanotubes in aqueous solutions with tRNA. Zaremba O, Goldt A, Ramirez-Morales M, Khabushev E, Shulga E, Eremin T, Prikazchikova T, Orekhov A, Grebenko A, Zatsepin T, Obraztsova E, Nasibulin A. Carbon, 2019. https://doi.org/10.1016/j.carbon.2019.05.076
- Synthesis of beta-diketone DNA derivatives for affinity modification of proteins. Monakhova M, Kubareva E, Romanova E, Semkina A, Naberezhnov D, Rao D, Zatsepin T, Oretskaya T. Russ. J. Bioorg. Chem., 2019 45(2), 144-154.
- Non-coding RNA in liver regeneration – from molecular mechanisms to clinical implications. Sergeeva O, Sviridov E, Zatsepin T. Seminars Liver Dis., 2019. 40(1):70-83. doi: 10.1055/s-0039-1693513
- Oligonucleotide Primers with G8AE-Clamp Modifications for RT-qPCR Detection of the Low-Copy dsRNA. Zatsepin T, Varizhuk A, Dedkov V, Shipulin G, Aralov G. Methods Mol Biol., 2019 1973, 281-297
- eIF4G2 balances its own mRNA translation via a PCBP2-based feedback loop. Smirnova V, Shestakova E, Bikmetov D, Chugunova A, Osterman I, Serebryakova M, Sergeeva O, Zatsepin T, Shatsky I, Terenin I. RNA, accepted 2019. doi:10.1261/rna.065623.118
- DNA i-motifs with guanidino-i-clamp residues: the counterplay between kinetics and thermodynamics and implications for the design of pH sensors. Tsvetkov V, Zatsepin T, Turaev A, Farzan V, Pozmogova G, Aralov A, Varizhuk A. Comput. J., Struct. Biotech. 2019, 17, , 527-536
-
Integrator is a key component of human telomerase RNA biogenesis. Rubtsova M, Vasilkova D, Moshareva M, Malyavko A, Meerson M, Zatsepin T, Naraykina Y, Beletsky AV, Ravin N, Dontsova O. Sci Rep. 2019 Feb 8;9(1):1701. doi: 10.1038/s41598-018-38297-6.
- Novel homo Yin-Yang probes improve sensitivity in RT-qPCR detection of low copy HIV RNA. Farzan V, Kvach M, Aparin I, Kireev D, Prikazchikova T, Ustinov A, Shmanai V, Shipulin G, Korshun V, Zatsepin T. Talanta. 01.03.2019;194:226-232. doi: 10.1016/j.talanta.2018.10.043.
- A study on endonuclease BspD6I and its stimulus-responsive switching by modified oligonucleotides L.A.Abrosimova, A.Y.Migur, E.A.Kubareva, T.S.Zatsepin, A.V.Gavshina, A.K.Yunusova, T.A.Perevyazova, A.Pingoud, T.S.Oretskaya. PLoS One, 13(11), e0207302 (2018).
- Novel Cluster and Monomer-Based GalNAc Structures Induce Effective Uptake of siRNAs in Vitro and in Vivo. Sharma V, Osborn M, Hassler M, Echeverria D, Ly S, Ulashchik E, Martynenko-Makaev Y, Shmanai V, Zatsepin T, Khvorova A, Watts JK. Bioconjug Chem. 2018 Jul 18;29(7):2478-2488. doi: 10.1021/acs.bioconjchem.8b00365. Epub 2018 Jul 2.
-
Structure and function of the N-terminal domain of the yeast telomerase reverse transcriptase. Petrova O, Mantsyzov A, Rodina E, Efimov S, Hackenberg C, Hakanpää J, Klochkov V, Lebedev A, Chugunova A, Malyavko A, Zatsepin T, Mishin A, Zvereva M, Lamzin V, Dontsova O, Polshakov V. Nucleic Acids Res. 2018 Feb 16;46(3):1525-1540. doi: 10.1093/nar/gkx1275.
-
i-Clamp phenoxazine for the fine tuning of DNA i-motif stability. Tsvetkov V, Zatsepin T, Belyaev E, Kostyukevich Y, Shpakovski G, Podgorsky V, Pozmogova G, Varizhuk A, Aralov A. Nucleic Acids Res. 2018, 10, 1093 121.
- Application of sorting and next generation sequencing to study 5΄-UTR influence on translation efficiency in Escherichia coli. Evfratov S, Osterman I, Komarova E, Pogorelskaya A, Rubtsova M, Zatsepin T, Semashko T, Kostryukova E, Mironov A, Burnaev E, Krymova E, Gelfand M, Govorun V, Bogdanov A, Sergiev P, Dontsova O. Nucleic Acids Res. 2017 Apr 7;45(6):3487-3502. doi: 10.1093/nar/gkw1141. PubMed PMID: 27899632; PubMed Central PMCID: PMC5389652.
- Automated Solid-Phase Click Synthesis of Oligonucleotide Conjugates: From Small Molecules to Diverse N-Acetylgalactosamine Clusters. Farzan V, Ulashchik E, Martynenko-Makaev Y, Kvach M, Aparin I, Brylev V, Prikazchikova T, Maklakova S, Majouga A, Ustinov A, Shipulin G, Shmanai V, Korshun V, Zatsepin T. Bioconjug Chem. 2017 Oct 18;28(10):2599-2607. doi: 10.1021/acs.bioconjchem.7b00462. Epub 2017 Oct 4.
- Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing Hao Yin, Chun-Qing Song, Sneha Suresh, Qiongqiong Wu, Walsh S, Hyunsik Rhym L, Mintzer E, Fatih Bolukbasi, Zhu J, Kauffman K, Haiwei Mou, Oberholzer A, Junmei Ding, Suet-Yan Kwan, Bogorad R, Zatsepin T, Koteliansky V, Wolfe S, Wen Xue, Langer R, Anderson C. Nature Biotechnology. 2017, 35, 1179–1187.
- Synthesis and biological evaluation of novel mono- and bivalent ASGP-R-targeted drug-conjugates Petrov R, Maklakova S, Ivanenkov Y, Petrov S, Sergeeva O, Yamansarov E, Saltykova I, Kireev I, Alieva I, Deyneka E, Sofronova A, Aladinskaia A, Trofimenko A, Yamidanov R, Kovalev S, Kotelianski V, Zatsepin T, Beloglazkina E, Majouga A. Bioorg Med Chem Lett. 2017, S0960-894X, (17), 31192-7.
- Synthesis and biological evaluation of novel doxorubicin-containing ASGP-R-targeted drug-conjugates. Y.A. Ivanenkov, A.G. Majouga, R.A. Petrov, S.A. Petrov, S.V. Kovalev, S.Y. Maklakova, E.Y. Yamansarov, I.V. Saltykova, E.V. Deyneka, G.I. Filkov, V.E. Kotelianski, T.S. Zatsepin, E.K. Beloglazkina. Bioorg. Med. Chem. Lett., 28(3), 503-508 (2018).
- Fine Tuning of Pyrene Excimer Fluorescence in Molecular Beacons by Alteration of the Monomer Structure Aparin I, Proskurin G, Golovin A, Ustinov A, Formanovsky A, Zatsepin T, Korshun V. J Org Chem. 2017, 6, 82, (19), 10015-10024.
- Automated solid phase click synthesis of oligonucleotide conjugates: from small molecules to diverse N-acetylgalactosamine clusters Farzan V, Ulashchik E, Martynenko-Makaev Y, Kvach M, Aparin I, Brylev V, Prikazchikova T, Maklakova S, Majouga A, Ustinov A, Shipulin G, Shmanai V, Korshun V, Zatsepin T. Bioconjug Chem. 2017, 18, 10, 1021
-
Human Ku70 protein binds hairpin RNA and double stranded DNA through two different sites. Anisenko A, Knyazhanskaya E, Zatsepin T, Gottikh M. Biochimie. 2017 Jan;132:85-93. doi: 10.1016/j.biochi.2016.11.001.
- Characterization of HIV-1 integrase interaction with human Ku70 protein and initial implications for drug targeting Anisenko A, Knyazhanskaya E, Zalevsky A, Agapkina J, Sizov A, Zatsepin T, Gottikh M. Sci Rep. 2017, 7, 1, 5649
- Design of 2?-phenylethynylpyrene excimer forming DNA/RNA probes for homogeneous SNP detection: The attachment manner matters Astakhova K, Golovin, Prokhorenkov I, Ustinov A, Stepanova I, Zatsepin T., Korshun V. Tetrahedron, 2017, 73, 3220–3230
- Oligonucleotide inhibitors of HIV-1 integrase efficiently inhibit HIV-1 reverse transcriptase. S.P. Korolev, T.S. Zatsepin, M.B. Gottikh. ,Russ. J. Bioorg. Chem., 43 (2), 135-139 (2017)
- Synthesis of oligonucleotides containing novel G-clamp analog with C8-tethered group in phenoxazine ring: implication to qPCR detection of the low-copy kemerovo virus dsRNA Varizhuk A, Zatsepin T, Golovin A, Belyaev E, Kostyukevich Y, Dedkov V, Shipulin G, Shpakovski G, Aralov A. . Bioorg. Med. Chem., 2017, 25, 3597-3605
- Specificity of SNP detection with molecular beacons is improved by stem and loop separation with spacers Farzan V, Markelov M, Skoblov A, Shipulin G, Zatsepin T. Analyst, 2017 142, 945-950.
- HumanKu70 protein binds hairpin RNA and double stranded DNA through two different sites. Anisenko A, Knyazhanskaya E, Zatsepin T, Gottikh M. Biochimie, 2017, 132, 85–93.
- lncRNA in the liver: Prospects for fundamental research and therapy by RNA interference. Smekalova E, Kotelevtsev Y, Leboeuf D, Shcherbinina E, Fefilova A, Zatsepin T, Koteliansky V. Biochimie. 2016. pii: S0300-9084(16)30116-X. doi: 10.1016/j.biochi.2016.06.007.
- Cy5/BHQ dye–quencher pairs in fluorogenic qPCR probes: effects of charge and hydrophobicity. Farzan V, Aparin I, Veselova O, Podkolzin A, Shipulin G, Korshun V, Zatsepin T. Anal. Methods, 2016, 8, 5826-5831
- Lipid nanoparticles for targeted siRNA delivery – going from bench to bedside. Zatsepin T, Kotelevtsev Y, Koteliansky V. Int J Nanomedicine. 2016 ;11:3077-86.
-
mRNA-Based Therapeutics – Advances and Perspectives.Sergeeva O, Koteliansky V, Zatsepin T. Biochemistry (Mosc). 2016Jul;81(7):709-22. doi: 10.1134/S0006297916070075.
- Tetrahedral DNA conjugates from pentaerythritol-based polyazides. Ponomarenko A, Brylev V, Sapozhnikova K, Ustinov A, Prokhorenko I, Zatsepin T, Korshun V Tetrahedron 2016 72: 2386–2391.
- 1-Phenylethynylpyrene (PEPy) as a novel blue-emitting dye for qPCR assay. Aparin I, Farzan V, Veselova O, Chistov A, Podkolzin A, Ustinov A, Shipulin G, Formanovsky A, Korshun V, Zatsepin T . Analyst. 2016 ;141(4):1331-8.
- Analysis of the cleavage mechanism by protein-only RNase P using precursor tRNA substrates with modifications at the cleavage site Walczyk D, Gößringer M, Rossmanith W, Zatsepin T, Oretskaya T, Hartmann R, J. Mol. Biol.2016, 428(24 Pt B), 4917-4928.
- Molecular beacons with JOE dye: Influence of linker and 3′ couple quencher. Tsybulsky D, Kvach M, Ryazantsev D, Aparin I, Stakheev A, Prokhorenko I, Martynenko Y, Gontarev S, Formanovsky A, Zatsepin T, Shmanai V, Korshun V, Zavriev S. Mol Cell Probes. 2016 Oct.
- Solid- and solution-phase synthesis and application of R6G dual-labeled oligonucleotide probes. Skoblov A, Vichuzhanin M, Farzan V, Veselova O, Konovalova T, Podkolzin A, Shipulin G, Zatsepin T. Bioorg Med Chem. 2015 Oct 15;23(20):6749-56. doi: 10.1016/j.bmc.2015.08.041. Epub 2015 Sep 1.
- TERRA mimicking ssRNAs prevail over the DNA substrate for telomerase in vitro due to interactions with the alternative binding site.Azhibek D, Skvortsov D, Andreeva A, Zatsepin T, Arutyunyan A, Zvereva M, Dontsova O. J Mol Recognit. 2015 Dec 16. doi: 10.1002/jmr.2521
- Oligonucleotide inhibitors of telomerase: prospects for anticancer therapy and diagnostics. Zvereva M, Zatsepin T, Azhibek D, Shubernetskaya O, Shpanchenko O, Dontsova O. Biochemistry (Mosc). 2015 Mar
- Chimeric bifunctional oligonucleotides as a novel tool to invade telomerase assembly Azhibek D, Zvereva M, Zatsepin T, Rubtsova M, Dontsova O. Nucleic Acids Res. 2014 Nov 1;42(15):9531-42. doi: 10.1093/nar/gku688. Epub 2014 Jul 31.
Current
Research grants/Industry funded research
Completed
Research grants/Industry funded research
Skoltech Biomedical Initiative
Next Generation Program: Skoltech – MIT Joint Projects
 |
“Targeted RNA delivery in vivo for therapy by RNAi and genome editing” and “Regulation of Antitumor Response via Modulation of Ubr‐Ubiquitin Ligases In Vivo” Prof. Daniel Anderson, Massachusetts Institute of Technology |
 |
“Novel RNA conjugates for targeted and functional RNA delivery” Prof. Vadim Shmanai, Laboratory of Bioconjugate Chemistry |
 |
“Mechanisms of hepatocyte polarization in vivo” Prof. Marino Zerial, Zerial Lab |
 |
”Cancer cell – specific RNA interference in vivo” Prof. Dr. Andriy Mokhir, Friedrich-Alexander-Universität Erlangen-Nürnberg |